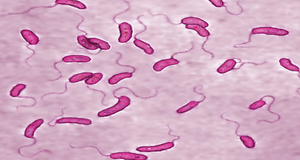

Vibrio cholerae is a highly motile, comma-shaped gram-negative bacteria with a single polar flagellum, belonging to the Vibrionaceae family, whose ecological niche is in salt or brackish water, usually associated with zooplankton and shellfish[1,2], and is transmitted through contaminated water and infects the host[3].
Cholera is a severe, life-threatening diarrheal disease that can kill its victims within hours if left untreated[4]. Until recently, cholera is still endemic in almost 69 countries, accounting for around 2.86 million cases and 95,000 deaths annually[5].
V. cholerae has been classified based on the surface somatic O antigens and more than 200 serogroups have been identified to date[6]. Of these serogroups, O1 and O139 have been associated with cholera epidemics, with O1 being further differentiated into two biotypes, classical (Cla) and El Tor (ET)[7,8]. There have been seven pandemics of cholera since 1817, The Cla biotype is believed to have caused the first six pandemics, whereas the ET biotype replaced Cla globally to cause the seventh cholera pandemic that has been ongoing since 1961[9]. O139 isolates were first identified in Bangladesh and India in 1992, which were found to be derived from the ET biotype and have not spread beyond Asia[10].
Samples collection date:
Samples host information:
Biotype information:
Serotype information:
Samples MLST information:
Samples phylogroup information:
Samples Virulence information:
Samples Resistance information:
[1]Clemens J D, Nair G B, Ahmed T, Qadri F, Holmgren J. Cholera. Lancet. 2017;390(10101):1539–1549.
[2]Kanungo S, Azman AS, Ramamurthy T, Deen J, Dutta S. Cholera. Lancet. 2022;399(10333):1429-1440.
[3]Yoon SH, Waters CM. Vibrio cholerae. Trends Microbiol. 2019;27(9):806-807.
[4]Harris JB, LaRocque RC, Qadri F, Ryan ET, Calderwood SB. Cholera. Lancet. 2012;379: 2466–2476.
[5]Somboonwit C, Menezes L J, Holt D A, Sinnott J T, Shapshak P. Current views and challenges on clinical cholera. Bioinformation. 2017;13(12):405–409.
[6]Chatterjee SN, Chaudhuri K. Lipopolysaccharides of Vibrio cholerae. I. Physical and chemical characterization. Biochim Biophys Acta. 2003;1639: 65–79.
[7]Kaper JB, Morris JG Jr, Levine MM. Cholera. Clin Microbiol Rev. 1995;8(1):48-86.
[8]Hraib M, Alaidi S, Jouni S, Saad S, Muna M, et al. Cholera: An Overview with Reference to the Syrian Outbreak. Avicenna J Med. 2023;13(4):199-205.
[9]Mutreja A, Kim DW, Thomson NR, Connor TR, Lee JH, et al. Evidence for several waves of global transmission in the seventh cholera pandemic. Nature. 2011;477: 462–465.
[10]Wang H, Yang C, Sun Z, Zheng W, Zhang W, et al. Genomic epidemiology of Vibrio cholerae reveals the regional and global spread of two epidemic non-toxigenic lineages. PLoS Negl Trop Dis. 2020;14(2):e0008046.